| Source.Country | Type | No..samples |
|---|---|---|
| Brazil | Year-long cohort study | 886 |
| Thailand | Year-long cohort study | 680 |
| Solomon Islands | Year-long cohort study | 709 |
| Volunteer Blood Donor Registry | Negative control | 98 |
| Australian Red Cross | Negative control | 97 |
| Thai Red Cross | Negative control | 69 |
| Brazil Blood Donor Registry | Negative control | 96 |
7 Classification
This tutorial focuses on understanding (i) what the classification algorithm aims to do, (ii) how to interpret these data, and (iii) how to visualise these data.
7.1 P. vivax Machine Learning Classification Algorithm
Plasma samples from year-long observational cohort studies conducted in malaria-endemic regions in Thailand (Kanchanaburi and Ratchaburi provinces), Brazil (Manaus) and Solomon Islands (Ngella) were measured for antigen-specific IgG antibody responses toward a panel of eight antigens using the method outlined in (Longley et al. 2020).
Enrolled individuals in the year-long cohort studies provided a blood sample every month. Light microscopy and qPCR targeting the blood-stages of P. vivax were performed to detect which individuals were infected and at which time point during these studies. IgG antibody responses towards a panel of P. vivax antigens were measured at the last visit of the year-long study, enabling us to characterise antibody responses related with time since P. vivax infection. Negative controls from the Australian Red Cross, Brazil Red Cross, Thai Red Cross and the Volunteer Blood Donor Registry in Victoria, Australia were included. These data were used as our training dataset (#tbl-table1).
This work develops a sero-diagnostic test to balance the selection of serological exposure markers that are associated with high classification performance, with the selection of proteins that are easier to manufacture and are more stable.
(#tbl-table2) outlines the top eight P. vivax proteins selected in the model (discussed further below in the random forest classification methods), their associated lifecycle stage and description of the proteins and expression system.
| Antigen | Gene.ID..PlasmoDB. |
|---|---|
| RBP2b | PVX_094255 |
| MSP1-19 | PVX_099980 |
| Pv-fam-a | PVX_096995 |
| MSP5 | PVX_003770 |
| EBP | KMZ83376.1 |
| PTEX150 | PVX_084720 |
| PvCSS | PVX_086200 |
| MSP8 | PVX_097625 |
Random forests are a machine learning algorithm which creates an “ensemble” or “sets” of decision trees (i.e., a “forest”) from slightly different versions of the training dataset. Each ensemble makes a separate prediction which is then averaged to get a more accurate and stable final prediction, resulting in good classification performance compared to a single decision tree and little overfitting to the training data due to the use of multiple trees.
We trained a random forest classification algorithm to learn the patterns in antibody response for someone infected in the previous nine months. Because of our sampling method, our training dataset includes “true positives” and “true negatives” which allows us to compare to the model predicted “positives” and “negatives”.
A random forest classification algorithm was created for all possible combinations for the eight antigens using 10-fold cross-validations with five repeats. The final random forest was fit with 1,000 trees and all hyperparameters were default.
7.1.1 Setup
library(SeroTrackR)
library(tidyverse)
your_raw_data <- c(
system.file("extdata", "example_MAGPIX_plate1.csv", package = "SeroTrackR"),
system.file("extdata", "example_MAGPIX_plate2.csv", package = "SeroTrackR"),
system.file("extdata", "example_MAGPIX_plate3.csv", package = "SeroTrackR")
)
your_plate_layout <- system.file("extdata", "example_platelayout_1.xlsx", package = "SeroTrackR")
sero_data <- readSeroData(
raw_data = your_raw_data,
platform = "magpix", # default
version = "4.2" # default
)
plate_list <- readPlateLayout(
plate_layout = your_plate_layout,
sero_data = sero_data
)
qc_results <- runQC(
sero_data = sero_data, # load in your serological data variable
plate_list = plate_list # load your plate list variable
)
mfi_to_rau_output <- MFItoRAU(
sero_data = sero_data,
plate_list = plate_list,
qc_results = qc_results,
std_point = 10
)The classifyResults() function classifies samples as recently exposed within the last nine months (“seropositive”) or not (“seronegative”).
7.2 Which algorithm do I choose?
The algorithm_type chosen can be “antibody_model” or “antibody_model_excLF016”.
antibody_model: PvSeroApp Algorithm: The PvSeroApp Model contains all top 8 antigens.
antibody_model_excLF016: As the name suggests, this model contains the antigens in PvSeroApp except for LF016.
7.3 Sensitivity and Specificity
7.3.1 Some definitions
- True Positive (TP), True Negative (TN), False Positive (FP), False Negative (FN).
- Sensitivity: Tests the proportion of people who test positive among all of those who actually have the disease.
- Sens = TP / (TP + FN).
- Specificity: Tests the proportion of people who test negative among all those who actually do not have the disease.
- Spec = TN / (TN + FP).
- Positive Predictive Value (PPV): Probability that following a positive test result, that individual will truly have the specific disease.
- PPV = TP / (TP + FP).
- Negative Predictive Value (NPV): Probability that following a negative test result, the individual will truly not have that specific disease.
- NPV = TN / (TN + FN).
7.3.2 Options
We have provided multiple options for you to choose which classification you would like to run. The sens_spec can be:
- “balanced” (default)
- “85% sensitivity”
- “90% sensitivity”
- “95% sensitivity”
- “85% specificity”
- “90% specificity”
- “95% specificity”
7.3.3 Which sensitivity/specificity threshold do I use?
The balanced sensitivity / specificity is when both metrics are at their highest and correspond to 81% each.
High sensitivity with lower specificity maximises the identification and treatment of all likely hypnozoite carriers at the cost of some over-treatment. As radical cure carries a risk of haemolysis in G6PD-deficient individuals10 and G6PD screening can add a substantial burden to healthcare costs, operational use of a high-sensitivity, low-specificity approach is unlikely.
High specificity with lower sensitivity is suited for surveillance settings and aims to correctly identify non-carriers and minimise treatment of uninfected individual, while the impact of false negatives can be mitigated by increasing survey size.
7.4 Applying the Random Forest
classifyResults_output <- classifyResults(
mfi_to_rau_output,
algorithm_type = "antibody_model",
sens_spec = "balanced",
qc_results
)The results from the classification are displayed in a table. The exposure status (seropositive/seronegative) is displayed for each sample.
To form an educated guess about the seropositivity/exposure status for your samples, take into consideration all relevant epidemiological information that you may have available to you (e.g. time since possible exposure, timing of sampling with respect to incidence of P. vivax infections/cases) and explore the results using the classifier/s.
classifyResults_output| SampleID | Plate | QC_total | PvEBP | Pv-fam-a | PvMSP5 | PvMSP1-19 | PvMSP8 | PvRBP2b | PvPTEX150 | PvCSS | pred_class_max |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ABC013 | plate1 | pass | 0.0008182 | 0.0013984 | 0.0002178 | 0.0001307 | 0.0000544 | 0.0005770 | 0.0008128 | 0.0002842 | seropositive |
| ABC097 | plate2 | pass | 0.0009351 | 0.0014014 | 0.0002126 | 0.0001217 | 0.0000447 | 0.0015043 | 0.0008057 | 0.0002839 | seropositive |
| ABC181 | plate3 | pass | 0.0009247 | 0.0014825 | 0.0002103 | 0.0001255 | 0.0000532 | 0.0003089 | 0.0008214 | 0.0002777 | seropositive |
| ABC022 | plate1 | pass | 0.0200000 | 0.0194207 | 0.0008140 | 0.0034202 | 0.0006996 | 0.0007638 | 0.0017080 | 0.0028056 | seropositive |
| ABC106 | plate2 | pass | 0.0200000 | 0.0166134 | 0.0008215 | 0.0033760 | 0.0006723 | 0.0200000 | 0.0016312 | 0.0027950 | seropositive |
| ABC190 | plate3 | pass | 0.0200000 | 0.0188708 | 0.0008173 | 0.0038346 | 0.0006613 | 0.0200000 | 0.0017157 | 0.0029083 | seropositive |
| ABC023 | plate1 | pass | 0.0002470 | 0.0066817 | 0.0001356 | 0.0001112 | 0.0001541 | 0.0008823 | 0.0103646 | 0.0010006 | seropositive |
| ABC107 | plate2 | pass | 0.0002679 | 0.0063880 | 0.0001253 | 0.0001013 | 0.0001431 | 0.0104717 | 0.0092600 | 0.0010103 | seropositive |
| ABC191 | plate3 | pass | 0.0002668 | 0.0071560 | 0.0001313 | 0.0001068 | 0.0001443 | 0.0200000 | 0.0106456 | 0.0010214 | seropositive |
| ABC024 | plate1 | pass | 0.0004662 | 0.0003515 | 0.0002854 | 0.0001167 | 0.0000897 | 0.0003179 | 0.0002972 | 0.0004277 | seropositive |
| ABC108 | plate2 | pass | 0.0005239 | 0.0003427 | 0.0002835 | 0.0001070 | 0.0000793 | 0.0002930 | 0.0003045 | 0.0004299 | seropositive |
| ABC192 | plate3 | pass | 0.0005171 | 0.0003445 | 0.0002764 | 0.0001120 | 0.0000853 | 0.0014281 | 0.0002999 | 0.0004233 | seropositive |
| ABC014 | plate1 | pass | 0.0000505 | 0.0003778 | 0.0000420 | 0.0000830 | 0.0000195 | 0.0002524 | 0.0001229 | 0.0001354 | seropositive |
| ABC098 | plate2 | pass | 0.0000478 | 0.0003692 | 0.0000263 | 0.0000718 | 0.0000195 | 0.0001479 | 0.0001221 | 0.0001330 | seronegative |
| ABC182 | plate3 | pass | 0.0000527 | 0.0003718 | 0.0000436 | 0.0000800 | 0.0000195 | 0.0000604 | 0.0001195 | 0.0001316 | seronegative |
| ABC015 | plate1 | pass | 0.0000672 | 0.0002221 | 0.0000724 | 0.0001473 | 0.0000657 | 0.0002511 | 0.0002782 | 0.0008855 | seropositive |
| ABC099 | plate2 | pass | 0.0000654 | 0.0002130 | 0.0000580 | 0.0001392 | 0.0000557 | 0.0001953 | 0.0002851 | 0.0008942 | seronegative |
| ABC183 | plate3 | pass | 0.0000700 | 0.0002132 | 0.0000719 | 0.0001416 | 0.0000635 | 0.0001722 | 0.0002804 | 0.0009003 | seronegative |
| ABC016 | plate1 | pass | 0.0000569 | 0.0002418 | 0.0000364 | 0.0000400 | 0.0000195 | 0.0003184 | 0.0001103 | 0.0001568 | seropositive |
| ABC100 | plate2 | pass | 0.0000545 | 0.0002326 | 0.0000208 | 0.0000274 | 0.0000195 | 0.0000470 | 0.0001083 | 0.0001546 | seronegative |
| ABC184 | plate3 | pass | 0.0000593 | 0.0002328 | 0.0000385 | 0.0000401 | 0.0000195 | 0.0000860 | 0.0001063 | 0.0001521 | seronegative |
| ABC017 | plate1 | pass | 0.0003416 | 0.0002480 | 0.0001016 | 0.0001933 | 0.0000623 | 0.0003606 | 0.0014239 | 0.0003288 | seropositive |
| ABC101 | plate2 | pass | 0.0003780 | 0.0002388 | 0.0000891 | 0.0001875 | 0.0000524 | 0.0003529 | 0.0013725 | 0.0003292 | seropositive |
| ABC185 | plate3 | pass | 0.0003741 | 0.0002390 | 0.0000992 | 0.0001870 | 0.0000604 | 0.0003426 | 0.0014322 | 0.0003225 | seropositive |
| ABC018 | plate1 | pass | 0.0000645 | 0.0003929 | 0.0000615 | 0.0000353 | 0.0000213 | 0.0000486 | 0.0002795 | 0.0001341 | seronegative |
| ABC102 | plate2 | pass | 0.0000625 | 0.0003844 | 0.0000466 | 0.0000227 | 0.0000195 | 0.0001498 | 0.0002864 | 0.0001317 | seronegative |
| ABC186 | plate3 | pass | 0.0000671 | 0.0003875 | 0.0000618 | 0.0000357 | 0.0000229 | 0.0001436 | 0.0002817 | 0.0001304 | seronegative |
| ABC019 | plate1 | pass | 0.0001412 | 0.0018592 | 0.0001734 | 0.0000645 | 0.0002753 | 0.0007980 | 0.0002325 | 0.0003450 | seropositive |
| ABC103 | plate2 | pass | 0.0001468 | 0.0018617 | 0.0001656 | 0.0000525 | 0.0002624 | 0.0010216 | 0.0002382 | 0.0003457 | seropositive |
| ABC187 | plate3 | pass | 0.0001492 | 0.0019939 | 0.0001674 | 0.0000627 | 0.0002566 | 0.0002739 | 0.0002334 | 0.0003389 | seropositive |
| ABC020 | plate1 | pass | 0.0000195 | 0.0000508 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000257 | 0.0000216 | seronegative |
| ABC104 | plate2 | pass | 0.0000195 | 0.0000463 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000199 | seronegative |
| ABC188 | plate3 | pass | 0.0000195 | 0.0000513 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000272 | seronegative |
| ABC021 | plate1 | pass | 0.0000195 | 0.0000457 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000269 | 0.0000216 | seronegative |
| ABC105 | plate2 | pass | 0.0000195 | 0.0000416 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000199 | seronegative |
| ABC189 | plate3 | pass | 0.0000195 | 0.0000469 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000272 | seronegative |
| ABC025 | plate1 | pass | 0.0000548 | 0.0004048 | 0.0003595 | 0.0000374 | 0.0000303 | 0.0001647 | 0.0005794 | 0.0001809 | seropositive |
| ABC109 | plate2 | pass | 0.0000523 | 0.0003965 | 0.0003607 | 0.0000249 | 0.0000219 | 0.0002329 | 0.0005830 | 0.0001790 | seropositive |
| ABC193 | plate3 | pass | 0.0000571 | 0.0003999 | 0.0003500 | 0.0000377 | 0.0000313 | 0.0000965 | 0.0005866 | 0.0001755 | seropositive |
| ABC034 | plate1 | pass | 0.0000195 | 0.0000643 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000391 | 0.0000396 | seronegative |
| ABC118 | plate2 | pass | 0.0000195 | 0.0000591 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000274 | 0.0000373 | seronegative |
| ABC202 | plate3 | pass | 0.0000195 | 0.0000633 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000313 | 0.0000430 | seronegative |
| ABC035 | plate1 | pass | 0.0000621 | 0.0005748 | 0.0000869 | 0.0001314 | 0.0012803 | 0.0007887 | 0.0007035 | 0.0016005 | seropositive |
| ABC119 | plate2 | pass | 0.0000600 | 0.0005686 | 0.0000735 | 0.0001225 | 0.0012195 | 0.0020972 | 0.0007021 | 0.0016108 | seropositive |
| ABC203 | plate3 | pass | 0.0000646 | 0.0005798 | 0.0000855 | 0.0001262 | 0.0012299 | 0.0008669 | 0.0007116 | 0.0016533 | seropositive |
| ABC036 | plate1 | pass | 0.0005558 | 0.0045156 | 0.0001381 | 0.0135998 | 0.0053044 | 0.0200000 | 0.0045939 | 0.0042145 | seropositive |
| ABC120 | plate2 | pass | 0.0006290 | 0.0044161 | 0.0001280 | 0.0120747 | 0.0048849 | 0.0081155 | 0.0041953 | 0.0041467 | seropositive |
| ABC204 | plate3 | pass | 0.0006206 | 0.0048924 | 0.0001337 | 0.0144136 | 0.0052812 | 0.0075679 | 0.0046200 | 0.0043363 | seropositive |
| ABC026 | plate1 | pass | 0.0003966 | 0.0027886 | 0.0003080 | 0.0000742 | 0.0000240 | 0.0005895 | 0.0001766 | 0.0001341 | seropositive |
| ABC110 | plate2 | pass | 0.0004423 | 0.0027742 | 0.0003071 | 0.0000626 | 0.0000195 | 0.0006318 | 0.0001797 | 0.0001317 | seropositive |
| ABC194 | plate3 | pass | 0.0004370 | 0.0030211 | 0.0002987 | 0.0000718 | 0.0000255 | 0.0003783 | 0.0001754 | 0.0001304 | seropositive |
| ABC027 | plate1 | pass | 0.0001226 | 0.0039278 | 0.0001132 | 0.0000609 | 0.0001133 | 0.0106539 | 0.0103523 | 0.0060087 | seropositive |
| ABC111 | plate2 | pass | 0.0001260 | 0.0038648 | 0.0001014 | 0.0000488 | 0.0001027 | 0.0104750 | 0.0092492 | 0.0058252 | seropositive |
| ABC195 | plate3 | pass | 0.0001289 | 0.0042619 | 0.0001101 | 0.0000594 | 0.0001068 | 0.0074520 | 0.0106324 | 0.0060911 | seropositive |
| ABC028 | plate1 | pass | 0.0001586 | 0.0016662 | 0.0001399 | 0.0000775 | 0.0005192 | 0.0014748 | 0.0021344 | 0.0004708 | seropositive |
| ABC112 | plate2 | pass | 0.0001665 | 0.0016695 | 0.0001300 | 0.0000660 | 0.0004994 | 0.0011805 | 0.0020160 | 0.0004737 | seropositive |
| ABC196 | plate3 | pass | 0.0001683 | 0.0017797 | 0.0001355 | 0.0000748 | 0.0004875 | 0.0012294 | 0.0021415 | 0.0004676 | seropositive |
| ABC029 | plate1 | pass | 0.0001787 | 0.0011494 | 0.0001119 | 0.0000707 | 0.0000234 | 0.0001333 | 0.0005192 | 0.0002905 | seropositive |
| ABC113 | plate2 | pass | 0.0001892 | 0.0011507 | 0.0001001 | 0.0000590 | 0.0000195 | 0.0001228 | 0.0005246 | 0.0002903 | seronegative |
| ABC197 | plate3 | pass | 0.0001904 | 0.0012069 | 0.0001089 | 0.0000685 | 0.0000249 | 0.0001182 | 0.0005258 | 0.0002840 | seropositive |
| ABC030 | plate1 | pass | 0.0002881 | 0.0005573 | 0.0000811 | 0.0000589 | 0.0009290 | 0.0005579 | 0.0047086 | 0.0093027 | seropositive |
| ABC114 | plate2 | pass | 0.0003156 | 0.0005508 | 0.0000673 | 0.0000467 | 0.0008899 | 0.0058959 | 0.0042961 | 0.0088044 | seropositive |
| ABC198 | plate3 | pass | 0.0003132 | 0.0005611 | 0.0000801 | 0.0000575 | 0.0008844 | 0.0064372 | 0.0047368 | 0.0091448 | seropositive |
| ABC031 | plate1 | pass | 0.0001734 | 0.0013669 | 0.0001067 | 0.0000879 | 0.0007773 | 0.0019346 | 0.0024283 | 0.0085605 | seropositive |
| ABC115 | plate2 | pass | 0.0001832 | 0.0013697 | 0.0000945 | 0.0000769 | 0.0007462 | 0.0005900 | 0.0022792 | 0.0081434 | seropositive |
| ABC199 | plate3 | pass | 0.0001845 | 0.0014476 | 0.0001040 | 0.0000847 | 0.0007365 | 0.0049079 | 0.0024354 | 0.0084741 | seropositive |
| ABC032 | plate1 | pass | 0.0002395 | 0.0200000 | 0.0001362 | 0.0032068 | 0.0042219 | 0.0200000 | 0.0055463 | 0.0070994 | seropositive |
| ABC116 | plate2 | pass | 0.0002592 | 0.0186125 | 0.0001260 | 0.0031746 | 0.0039078 | 0.0108335 | 0.0050315 | 0.0068252 | seropositive |
| ABC200 | plate3 | pass | 0.0002583 | 0.0200000 | 0.0001319 | 0.0035905 | 0.0041862 | 0.0200000 | 0.0055933 | 0.0071250 | seropositive |
| ABC033 | plate1 | pass | 0.0000491 | 0.0002513 | 0.0000261 | 0.0000291 | 0.0000195 | 0.0000822 | 0.0001993 | 0.0000748 | seronegative |
| ABC117 | plate2 | pass | 0.0000463 | 0.0002421 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000754 | 0.0002037 | 0.0000722 | seronegative |
| ABC201 | plate3 | pass | 0.0000512 | 0.0002423 | 0.0000289 | 0.0000301 | 0.0000195 | 0.0000741 | 0.0001991 | 0.0000748 | seronegative |
| ABC037 | plate1 | pass | 0.0000913 | 0.0013441 | 0.0000420 | 0.0000565 | 0.0000503 | 0.0002643 | 0.0007898 | 0.0001276 | seropositive |
| ABC121 | plate2 | pass | 0.0000914 | 0.0013469 | 0.0000263 | 0.0000442 | 0.0000408 | 0.0001616 | 0.0007840 | 0.0001252 | seronegative |
| ABC205 | plate3 | pass | 0.0000953 | 0.0014224 | 0.0000436 | 0.0000553 | 0.0000495 | 0.0001548 | 0.0007983 | 0.0001242 | seronegative |
| ABC046 | plate1 | pass | 0.0002422 | 0.0005543 | 0.0000832 | 0.0000738 | 0.0004278 | 0.0004831 | 0.0012861 | 0.0003226 | seropositive |
| ABC130 | plate2 | pass | 0.0002623 | 0.0005478 | 0.0000695 | 0.0000622 | 0.0004112 | 0.0002307 | 0.0012462 | 0.0003230 | seropositive |
| ABC214 | plate3 | pass | 0.0002613 | 0.0005579 | 0.0000820 | 0.0000714 | 0.0004005 | 0.0005918 | 0.0012947 | 0.0003163 | seropositive |
| ABC047 | plate1 | pass | 0.0001511 | 0.0004119 | 0.0000450 | 0.0000305 | 0.0000195 | 0.0002178 | 0.0001802 | 0.0000923 | seropositive |
| ABC131 | plate2 | pass | 0.0001580 | 0.0004036 | 0.0000295 | 0.0000195 | 0.0000195 | 0.0000684 | 0.0001835 | 0.0000897 | seronegative |
| ABC215 | plate3 | pass | 0.0001600 | 0.0004073 | 0.0000465 | 0.0000313 | 0.0000195 | 0.0001315 | 0.0001793 | 0.0000910 | seronegative |
| ABC048 | plate1 | pass | 0.0000739 | 0.0010297 | 0.0000490 | 0.0000750 | 0.0000608 | 0.0005895 | 0.0004570 | 0.0005160 | seropositive |
| ABC132 | plate2 | pass | 0.0000725 | 0.0010298 | 0.0000335 | 0.0000634 | 0.0000510 | 0.0001044 | 0.0004638 | 0.0005197 | seronegative |
| ABC216 | plate3 | pass | 0.0000769 | 0.0010750 | 0.0000501 | 0.0000725 | 0.0000591 | 0.0004973 | 0.0004628 | 0.0005142 | seropositive |
| ABC038 | plate1 | pass | 0.0000195 | 0.0000508 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000269 | 0.0000216 | seronegative |
| ABC122 | plate2 | pass | 0.0000195 | 0.0000463 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000199 | seronegative |
| ABC206 | plate3 | pass | 0.0000195 | 0.0000513 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000195 | 0.0000272 | seronegative |
| ABC039 | plate1 | pass | 0.0001304 | 0.0015212 | 0.0001102 | 0.0004159 | 0.0002822 | 0.0005618 | 0.0002838 | 0.0002031 | seropositive |
| ABC123 | plate2 | pass | 0.0001346 | 0.0015245 | 0.0000982 | 0.0004202 | 0.0002692 | 0.0005078 | 0.0002909 | 0.0002015 | seropositive |
| ABC207 | plate3 | pass | 0.0001373 | 0.0016187 | 0.0001073 | 0.0004162 | 0.0002631 | 0.0009869 | 0.0002862 | 0.0001972 | seropositive |
| ABC040 | plate1 | pass | 0.0000765 | 0.0004950 | 0.0000766 | 0.0000807 | 0.0000270 | 0.0001100 | 0.0003782 | 0.0002318 | seropositive |
| ABC124 | plate2 | pass | 0.0000754 | 0.0004877 | 0.0000625 | 0.0000694 | 0.0000195 | 0.0002076 | 0.0003859 | 0.0002307 | seropositive |
| ABC208 | plate3 | pass | 0.0000797 | 0.0004948 | 0.0000758 | 0.0000779 | 0.0000283 | 0.0001436 | 0.0003827 | 0.0002255 | seronegative |
| ABC041 | plate1 | pass | 0.0001207 | 0.0003185 | 0.0000584 | 0.0000396 | 0.0000201 | 0.0001111 | 0.0001957 | 0.0002412 | seronegative |
| ABC125 | plate2 | pass | 0.0001238 | 0.0003095 | 0.0000433 | 0.0000270 | 0.0000195 | 0.0000850 | 0.0001999 | 0.0002402 | seronegative |
| ABC209 | plate3 | pass | 0.0001268 | 0.0003105 | 0.0000589 | 0.0000397 | 0.0000218 | 0.0001053 | 0.0001953 | 0.0002348 | seronegative |
| ABC042 | plate1 | pass | 0.0003817 | 0.0061971 | 0.0000940 | 0.0000956 | 0.0000413 | 0.0011964 | 0.0015070 | 0.0012158 | seropositive |
| ABC126 | plate2 | pass | 0.0004249 | 0.0059545 | 0.0000810 | 0.0000849 | 0.0000322 | 0.0012398 | 0.0014485 | 0.0012266 | seropositive |
| ABC210 | plate3 | pass | 0.0004200 | 0.0066577 | 0.0000921 | 0.0000919 | 0.0000413 | 0.0003908 | 0.0015152 | 0.0012481 | seropositive |
| ABC043 | plate1 | pass | 0.0001106 | 0.0003947 | 0.0000464 | 0.0000571 | 0.0000237 | 0.0004852 | 0.0012942 | 0.0003878 | seropositive |
| ABC127 | plate2 | pass | 0.0001125 | 0.0003862 | 0.0000309 | 0.0000449 | 0.0000195 | 0.0002608 | 0.0012536 | 0.0003893 | seropositive |
| ABC211 | plate3 | pass | 0.0001159 | 0.0003893 | 0.0000478 | 0.0000558 | 0.0000252 | 0.0002220 | 0.0013027 | 0.0003825 | seropositive |
| ABC044 | plate1 | pass | 0.0001622 | 0.0027317 | 0.0001337 | 0.0023118 | 0.0011158 | 0.0044115 | 0.0005151 | 0.0056849 | seropositive |
| ABC128 | plate2 | pass | 0.0001706 | 0.0027189 | 0.0001233 | 0.0023177 | 0.0010656 | 0.0046875 | 0.0005206 | 0.0055255 | seropositive |
| ABC212 | plate3 | pass | 0.0001722 | 0.0029585 | 0.0001295 | 0.0025618 | 0.0010676 | 0.0051104 | 0.0005216 | 0.0057795 | seropositive |
| ABC045 | plate1 | pass | 0.0051627 | 0.0008018 | 0.0001124 | 0.0000711 | 0.0002263 | 0.0005912 | 0.0003846 | 0.0003247 | seropositive |
| ABC129 | plate2 | pass | 0.0053468 | 0.0007990 | 0.0001006 | 0.0000594 | 0.0002143 | 0.0003546 | 0.0003922 | 0.0003251 | seropositive |
| ABC213 | plate3 | pass | 0.0056594 | 0.0008252 | 0.0001094 | 0.0000689 | 0.0002110 | 0.0003253 | 0.0003892 | 0.0003184 | seropositive |
| ABC049 | plate1 | pass | 0.0001153 | 0.0005906 | 0.0001234 | 0.0050223 | 0.0006990 | 0.0038292 | 0.0014106 | 0.0094299 | seropositive |
| ABC133 | plate2 | pass | 0.0001178 | 0.0005846 | 0.0001123 | 0.0048557 | 0.0006717 | 0.0041471 | 0.0013604 | 0.0089171 | seropositive |
| ABC217 | plate3 | pass | 0.0001210 | 0.0005967 | 0.0001198 | 0.0056405 | 0.0006606 | 0.0045122 | 0.0014189 | 0.0092588 | seropositive |
| ABC058 | plate1 | pass | 0.0003118 | 0.0023965 | 0.0004721 | 0.0001181 | 0.0039125 | 0.0200000 | 0.0033824 | 0.0200000 | seropositive |
| ABC142 | plate2 | pass | 0.0003431 | 0.0023918 | 0.0004765 | 0.0001085 | 0.0036279 | 0.0129060 | 0.0031273 | 0.0200000 | seropositive |
| ABC226 | plate3 | pass | 0.0003401 | 0.0025890 | 0.0004635 | 0.0001134 | 0.0038732 | 0.0137702 | 0.0033933 | 0.0200000 | seropositive |
| ABC059 | plate1 | pass | 0.0001675 | 0.0047330 | 0.0011415 | 0.0000559 | 0.0000570 | 0.0038731 | 0.0159424 | 0.0028020 | seropositive |
| ABC143 | plate2 | pass | 0.0001766 | 0.0046182 | 0.0011450 | 0.0000436 | 0.0000472 | 0.0016835 | 0.0142002 | 0.0027915 | seropositive |
| ABC227 | plate3 | pass | 0.0001781 | 0.0051239 | 0.0011645 | 0.0000547 | 0.0000556 | 0.0005000 | 0.0167561 | 0.0029046 | seropositive |
| ABC060 | plate1 | pass | 0.0002715 | 0.0017415 | 0.0002431 | 0.0002202 | 0.0000369 | 0.0004456 | 0.0003913 | 0.0001702 | seropositive |
| ABC144 | plate2 | pass | 0.0002962 | 0.0017445 | 0.0002392 | 0.0002157 | 0.0000280 | 0.0002076 | 0.0003989 | 0.0001681 | seropositive |
| ABC228 | plate3 | pass | 0.0002944 | 0.0018632 | 0.0002349 | 0.0002138 | 0.0000373 | 0.0002719 | 0.0003960 | 0.0001651 | seropositive |
| ABC050 | plate1 | pass | 0.0001815 | 0.0098569 | 0.0001462 | 0.0000561 | 0.0000425 | 0.0011679 | 0.0018457 | 0.0014488 | seropositive |
| ABC134 | plate2 | pass | 0.0001925 | 0.0091316 | 0.0001366 | 0.0000438 | 0.0000333 | 0.0002261 | 0.0017559 | 0.0014596 | seropositive |
| ABC218 | plate3 | pass | 0.0001935 | 0.0103116 | 0.0001414 | 0.0000549 | 0.0000424 | 0.0014854 | 0.0018531 | 0.0014936 | seropositive |
| ABC051 | plate1 | pass | 0.0001660 | 0.0003438 | 0.0000540 | 0.0000637 | 0.0000273 | 0.0003231 | 0.0018735 | 0.0002268 | seropositive |
| ABC135 | plate2 | pass | 0.0001748 | 0.0003350 | 0.0000387 | 0.0000516 | 0.0000195 | 0.0001528 | 0.0017810 | 0.0002256 | seropositive |
| ABC219 | plate3 | pass | 0.0001763 | 0.0003366 | 0.0000548 | 0.0000619 | 0.0000286 | 0.0001996 | 0.0018809 | 0.0002206 | seropositive |
| ABC052 | plate1 | pass | 0.0000907 | 0.0003680 | 0.0001310 | 0.0000400 | 0.0000429 | 0.0001532 | 0.0002825 | 0.0001903 | seronegative |
| ABC136 | plate2 | pass | 0.0000907 | 0.0003593 | 0.0001205 | 0.0000274 | 0.0000337 | 0.0001663 | 0.0002895 | 0.0001885 | seronegative |
| ABC220 | plate3 | pass | 0.0000947 | 0.0003616 | 0.0001270 | 0.0000401 | 0.0000428 | 0.0001593 | 0.0002848 | 0.0001847 | seronegative |
| ABC053 | plate1 | pass | 0.0002729 | 0.0017433 | 0.0001452 | 0.0000738 | 0.0001461 | 0.0005439 | 0.0006263 | 0.0019413 | seropositive |
| ABC137 | plate2 | pass | 0.0002978 | 0.0017463 | 0.0001356 | 0.0000622 | 0.0001351 | 0.0004619 | 0.0006282 | 0.0019485 | seropositive |
| ABC221 | plate3 | pass | 0.0002959 | 0.0018652 | 0.0001405 | 0.0000714 | 0.0001369 | 0.0010050 | 0.0006339 | 0.0020108 | seropositive |
| ABC054 | plate1 | pass | 0.0001427 | 0.0016726 | 0.0000651 | 0.0045487 | 0.0090314 | 0.0200000 | 0.0013554 | 0.0011121 | seropositive |
| ABC138 | plate2 | pass | 0.0001485 | 0.0016758 | 0.0000504 | 0.0044238 | 0.0082439 | 0.0200000 | 0.0013098 | 0.0011225 | seropositive |
| ABC222 | plate3 | pass | 0.0001508 | 0.0017867 | 0.0000652 | 0.0051118 | 0.0090366 | 0.0200000 | 0.0013639 | 0.0011388 | seropositive |
| ABC055 | plate1 | pass | 0.0020975 | 0.0011662 | 0.0006138 | 0.0000919 | 0.0016241 | 0.0014340 | 0.0051925 | 0.0022533 | seropositive |
| ABC139 | plate2 | pass | 0.0023613 | 0.0011676 | 0.0006206 | 0.0000810 | 0.0015390 | 0.0031830 | 0.0047211 | 0.0022558 | seropositive |
| ABC223 | plate3 | pass | 0.0023881 | 0.0012254 | 0.0006087 | 0.0000884 | 0.0015709 | 0.0034396 | 0.0052308 | 0.0023366 | seropositive |
| ABC056 | plate1 | pass | 0.0200000 | 0.0200000 | 0.0006379 | 0.0070091 | 0.0028312 | 0.0200000 | 0.0045887 | 0.0033729 | seropositive |
| ABC140 | plate2 | pass | 0.0200000 | 0.0200000 | 0.0006450 | 0.0066224 | 0.0026465 | 0.0200000 | 0.0041906 | 0.0033432 | seropositive |
| ABC224 | plate3 | pass | 0.0200000 | 0.0200000 | 0.0006337 | 0.0078070 | 0.0027814 | 0.0176802 | 0.0046146 | 0.0034887 | seropositive |
| ABC057 | plate1 | pass | 0.0001914 | 0.0200000 | 0.0001245 | 0.0000950 | 0.0001541 | 0.0200000 | 0.0088624 | 0.0013703 | seropositive |
| ABC141 | plate2 | pass | 0.0002038 | 0.0200000 | 0.0001135 | 0.0000843 | 0.0001431 | 0.0192940 | 0.0079396 | 0.0013813 | seropositive |
| ABC225 | plate3 | pass | 0.0002045 | 0.0200000 | 0.0001208 | 0.0000914 | 0.0001443 | 0.0151456 | 0.0090475 | 0.0014110 | seropositive |
| ABC061 | plate1 | pass | 0.0002976 | 0.0113481 | 0.0001672 | 0.0000792 | 0.0003708 | 0.0200000 | 0.0200000 | 0.0128668 | seropositive |
| ABC145 | plate2 | pass | 0.0003266 | 0.0103687 | 0.0001590 | 0.0000678 | 0.0003558 | 0.0074739 | 0.0196920 | 0.0119063 | seropositive |
| ABC229 | plate3 | pass | 0.0003240 | 0.0117330 | 0.0001615 | 0.0000765 | 0.0003465 | 0.0057903 | 0.0200000 | 0.0122411 | seropositive |
| ABC070 | plate1 | pass | 0.0005433 | 0.0182223 | 0.0001544 | 0.0165283 | 0.0130777 | 0.0200000 | 0.0058155 | 0.0053394 | seropositive |
| ABC154 | plate2 | pass | 0.0006143 | 0.0157285 | 0.0001453 | 0.0143388 | 0.0119087 | 0.0200000 | 0.0052676 | 0.0052041 | seropositive |
| ABC238 | plate3 | pass | 0.0006061 | 0.0178640 | 0.0001492 | 0.0170964 | 0.0130660 | 0.0067501 | 0.0058699 | 0.0054445 | seropositive |
| ABC071 | plate1 | pass | 0.0000875 | 0.0014835 | 0.0000585 | 0.0000545 | 0.0000463 | 0.0001220 | 0.0006337 | 0.0001629 | seronegative |
| ABC155 | plate2 | pass | 0.0000872 | 0.0014867 | 0.0000434 | 0.0000422 | 0.0000369 | 0.0001120 | 0.0006353 | 0.0001608 | seropositive |
| ABC239 | plate3 | pass | 0.0000913 | 0.0015769 | 0.0000590 | 0.0000534 | 0.0000458 | 0.0001080 | 0.0006414 | 0.0001580 | seronegative |
| ABC072 | plate1 | pass | 0.0001589 | 0.0038301 | 0.0001296 | 0.0013423 | 0.0028364 | 0.0145496 | 0.0037829 | 0.0026963 | seropositive |
| ABC156 | plate2 | pass | 0.0001668 | 0.0037724 | 0.0001190 | 0.0013636 | 0.0026512 | 0.0016456 | 0.0034810 | 0.0026887 | seropositive |
| ABC240 | plate3 | pass | 0.0001686 | 0.0041565 | 0.0001257 | 0.0014467 | 0.0027866 | 0.0029278 | 0.0037974 | 0.0027956 | seropositive |
| ABC062 | plate1 | pass | 0.0002459 | 0.0017333 | 0.0001536 | 0.0000904 | 0.0059064 | 0.0050882 | 0.0038324 | 0.0007895 | seropositive |
| ABC146 | plate2 | pass | 0.0002666 | 0.0017364 | 0.0001445 | 0.0000794 | 0.0054275 | 0.0034437 | 0.0035247 | 0.0007971 | seropositive |
| ABC230 | plate3 | pass | 0.0002655 | 0.0018541 | 0.0001485 | 0.0000870 | 0.0058899 | 0.0188942 | 0.0038475 | 0.0007994 | seropositive |
| ABC063 | plate1 | pass | 0.0001342 | 0.0008982 | 0.0002953 | 0.0000581 | 0.0000240 | 0.0001383 | 0.0019994 | 0.0010736 | seropositive |
| ABC147 | plate2 | pass | 0.0001389 | 0.0008967 | 0.0002938 | 0.0000459 | 0.0000195 | 0.0001277 | 0.0018946 | 0.0010838 | seropositive |
| ABC231 | plate3 | pass | 0.0001415 | 0.0009305 | 0.0002862 | 0.0000568 | 0.0000255 | 0.0001228 | 0.0020067 | 0.0010982 | seropositive |
| ABC064 | plate1 | pass | 0.0003349 | 0.0005816 | 0.0001199 | 0.0000826 | 0.0034099 | 0.0200000 | 0.0057515 | 0.0200000 | seropositive |
| ABC148 | plate2 | pass | 0.0003701 | 0.0005755 | 0.0001086 | 0.0000714 | 0.0031725 | 0.0005155 | 0.0052115 | 0.0200000 | seropositive |
| ABC232 | plate3 | pass | 0.0003664 | 0.0005871 | 0.0001165 | 0.0000797 | 0.0033653 | 0.0143496 | 0.0058041 | 0.0200000 | seropositive |
| ABC065 | plate1 | pass | 0.0002917 | 0.0021286 | 0.0002089 | 0.0001526 | 0.0035616 | 0.0200000 | 0.0066382 | 0.0200000 | seropositive |
| ABC149 | plate2 | pass | 0.0003198 | 0.0021284 | 0.0002031 | 0.0001447 | 0.0033101 | 0.0124392 | 0.0059889 | 0.0200000 | seropositive |
| ABC233 | plate3 | pass | 0.0003173 | 0.0022926 | 0.0002016 | 0.0001468 | 0.0035185 | 0.0050142 | 0.0067194 | 0.0200000 | seropositive |
| ABC066 | plate1 | pass | 0.0003899 | 0.0200000 | 0.0004239 | 0.0052843 | 0.0062343 | 0.0200000 | 0.0053963 | 0.0092426 | seropositive |
| ABC150 | plate2 | pass | 0.0004345 | 0.0200000 | 0.0004271 | 0.0050927 | 0.0057228 | 0.0200000 | 0.0048999 | 0.0087510 | seropositive |
| ABC234 | plate3 | pass | 0.0004294 | 0.0200000 | 0.0004147 | 0.0059310 | 0.0062211 | 0.0122844 | 0.0054394 | 0.0090909 | seropositive |
| ABC067 | plate1 | pass | 0.0001447 | 0.0003814 | 0.0000609 | 0.0000467 | 0.0000195 | 0.0002600 | 0.0005819 | 0.0004287 | seropositive |
| ABC151 | plate2 | pass | 0.0001508 | 0.0003728 | 0.0000459 | 0.0000342 | 0.0000195 | 0.0003227 | 0.0005854 | 0.0004309 | seropositive |
| ABC235 | plate3 | pass | 0.0001530 | 0.0003755 | 0.0000612 | 0.0000462 | 0.0000198 | 0.0003124 | 0.0005891 | 0.0004243 | seropositive |
| ABC068 | plate1 | pass | 0.0000601 | 0.0002882 | 0.0008196 | 0.0000469 | 0.0000195 | 0.0001680 | 0.0006037 | 0.0005286 | seropositive |
| ABC152 | plate2 | pass | 0.0000578 | 0.0002791 | 0.0008271 | 0.0000344 | 0.0000195 | 0.0002602 | 0.0006065 | 0.0005325 | seropositive |
| ABC236 | plate3 | pass | 0.0000625 | 0.0002796 | 0.0008232 | 0.0000464 | 0.0000195 | 0.0002504 | 0.0006111 | 0.0005273 | seropositive |
| ABC069 | plate1 | pass | 0.0001465 | 0.0006831 | 0.0001324 | 0.0007343 | 0.0041172 | 0.0200000 | 0.0116271 | 0.0086296 | seropositive |
| ABC153 | plate2 | pass | 0.0001527 | 0.0006785 | 0.0001220 | 0.0007486 | 0.0038132 | 0.0143344 | 0.0103725 | 0.0082052 | seropositive |
| ABC237 | plate3 | pass | 0.0001549 | 0.0006963 | 0.0001283 | 0.0007615 | 0.0040803 | 0.0083445 | 0.0120045 | 0.0085370 | seropositive |
| ABC073 | plate1 | pass | 0.0002083 | 0.0028578 | 0.0001660 | 0.0085473 | 0.0007000 | 0.0062469 | 0.0003337 | 0.0011927 | seropositive |
| ABC157 | plate2 | pass | 0.0002231 | 0.0028413 | 0.0001577 | 0.0079464 | 0.0006727 | 0.0004457 | 0.0003413 | 0.0012034 | seropositive |
| ABC241 | plate3 | pass | 0.0002233 | 0.0030971 | 0.0001603 | 0.0094265 | 0.0006616 | 0.0018792 | 0.0003373 | 0.0012238 | seropositive |
| ABC082 | plate1 | pass | 0.0000625 | 0.0006420 | 0.0000468 | 0.0000834 | 0.0000408 | 0.0002136 | 0.0002773 | 0.0003170 | seropositive |
| ABC166 | plate2 | pass | 0.0000603 | 0.0006368 | 0.0000312 | 0.0000722 | 0.0000317 | 0.0001693 | 0.0002842 | 0.0003172 | seropositive |
| ABC250 | plate3 | pass | 0.0000650 | 0.0006519 | 0.0000481 | 0.0000804 | 0.0000409 | 0.0001077 | 0.0002795 | 0.0003106 | seronegative |
| ABC083 | plate1 | pass | 0.0000666 | 0.0002854 | 0.0000714 | 0.0017211 | 0.0007687 | 0.0017111 | 0.0006387 | 0.0025842 | seropositive |
| ABC167 | plate2 | pass | 0.0000647 | 0.0002763 | 0.0000570 | 0.0017399 | 0.0007381 | 0.0019436 | 0.0006400 | 0.0025795 | seropositive |
| ABC251 | plate3 | pass | 0.0000692 | 0.0002768 | 0.0000710 | 0.0018811 | 0.0007282 | 0.0003513 | 0.0006463 | 0.0026798 | seropositive |
| ABC084 | plate1 | pass | 0.0000893 | 0.0039623 | 0.0000860 | 0.0000429 | 0.0000207 | 0.0005500 | 0.0008326 | 0.0007107 | seropositive |
| ABC168 | plate2 | pass | 0.0000892 | 0.0038974 | 0.0000724 | 0.0000304 | 0.0000195 | 0.0003840 | 0.0008244 | 0.0007174 | seropositive |
| ABC252 | plate3 | pass | 0.0000931 | 0.0042992 | 0.0000846 | 0.0000428 | 0.0000224 | 0.0002914 | 0.0008412 | 0.0007169 | seropositive |
| ABC074 | plate1 | pass | 0.0000872 | 0.0005436 | 0.0002240 | 0.0000613 | 0.0000240 | 0.0001853 | 0.0017859 | 0.0014200 | seropositive |
| ABC158 | plate2 | pass | 0.0000869 | 0.0005370 | 0.0002190 | 0.0000492 | 0.0000195 | 0.0005677 | 0.0017018 | 0.0014310 | seropositive |
| ABC242 | plate3 | pass | 0.0000909 | 0.0005465 | 0.0002163 | 0.0000597 | 0.0000255 | 0.0007772 | 0.0017935 | 0.0014634 | seropositive |
| ABC075 | plate1 | pass | 0.0004190 | 0.0010136 | 0.0001268 | 0.0000660 | 0.0000502 | 0.0005461 | 0.0013277 | 0.0016624 | seropositive |
| ABC159 | plate2 | pass | 0.0004686 | 0.0010135 | 0.0001160 | 0.0000541 | 0.0000407 | 0.0002542 | 0.0012844 | 0.0016723 | seropositive |
| ABC243 | plate3 | pass | 0.0004628 | 0.0010572 | 0.0001230 | 0.0000641 | 0.0000494 | 0.0007663 | 0.0013362 | 0.0017184 | seropositive |
| ABC076 | plate1 | pass | 0.0001548 | 0.0076392 | 0.0000821 | 0.0000742 | 0.0000541 | 0.0083323 | 0.0014196 | 0.0010793 | seropositive |
| ABC160 | plate2 | pass | 0.0001621 | 0.0072323 | 0.0000683 | 0.0000626 | 0.0000444 | 0.0005094 | 0.0013686 | 0.0010895 | seropositive |
| ABC244 | plate3 | pass | 0.0001640 | 0.0081271 | 0.0000810 | 0.0000718 | 0.0000529 | 0.0011934 | 0.0014279 | 0.0011043 | seropositive |
| ABC077 | plate1 | pass | 0.0001139 | 0.0005961 | 0.0000684 | 0.0000525 | 0.0000818 | 0.0004224 | 0.0017615 | 0.0021118 | seropositive |
| ABC161 | plate2 | pass | 0.0001162 | 0.0005902 | 0.0000538 | 0.0000401 | 0.0000715 | 0.0001649 | 0.0016797 | 0.0021167 | seropositive |
| ABC245 | plate3 | pass | 0.0001195 | 0.0006026 | 0.0000682 | 0.0000515 | 0.0000781 | 0.0001075 | 0.0017691 | 0.0021891 | seropositive |
| ABC078 | plate1 | pass | 0.0003381 | 0.0200000 | 0.0002190 | 0.0064864 | 0.0090958 | 0.0200000 | 0.0031907 | 0.0144574 | seropositive |
| ABC162 | plate2 | pass | 0.0003739 | 0.0200000 | 0.0002139 | 0.0061642 | 0.0083020 | 0.0200000 | 0.0029575 | 0.0132561 | seropositive |
| ABC246 | plate3 | pass | 0.0003701 | 0.0200000 | 0.0002115 | 0.0072453 | 0.0091011 | 0.0026367 | 0.0032003 | 0.0135633 | seropositive |
| ABC079 | plate1 | pass | 0.0065998 | 0.0007549 | 0.0000876 | 0.0000800 | 0.0006687 | 0.0023746 | 0.0005283 | 0.0003206 | seropositive |
| ABC163 | plate2 | pass | 0.0065843 | 0.0007514 | 0.0000741 | 0.0000686 | 0.0006428 | 0.0002533 | 0.0005335 | 0.0003209 | seropositive |
| ABC247 | plate3 | pass | 0.0070873 | 0.0007742 | 0.0000861 | 0.0000772 | 0.0006314 | 0.0005818 | 0.0005350 | 0.0003143 | seropositive |
| ABC080 | plate1 | pass | 0.0002327 | 0.0007571 | 0.0001062 | 0.0001170 | 0.0039092 | 0.0132681 | 0.0027010 | 0.0007148 | seropositive |
| ABC164 | plate2 | pass | 0.0002513 | 0.0007535 | 0.0000940 | 0.0001074 | 0.0036249 | 0.0004260 | 0.0025225 | 0.0007215 | seropositive |
| ABC248 | plate3 | pass | 0.0002507 | 0.0007765 | 0.0001036 | 0.0001124 | 0.0038699 | 0.0004817 | 0.0027085 | 0.0007211 | seropositive |
| ABC081 | plate1 | pass | 0.0002270 | 0.0011262 | 0.0001016 | 0.0001039 | 0.0000288 | 0.0003823 | 0.0003664 | 0.0001803 | seropositive |
| ABC165 | plate2 | pass | 0.0002447 | 0.0011274 | 0.0000891 | 0.0000936 | 0.0000205 | 0.0004457 | 0.0003741 | 0.0001784 | seropositive |
| ABC249 | plate3 | pass | 0.0002443 | 0.0011814 | 0.0000992 | 0.0000998 | 0.0000299 | 0.0000975 | 0.0003707 | 0.0001749 | seronegative |
| ABC085 | plate1 | pass | 0.0001581 | 0.0002036 | 0.0000378 | 0.0000573 | 0.0000201 | 0.0008026 | 0.0014746 | 0.0001543 | seropositive |
| ABC169 | plate2 | pass | 0.0001658 | 0.0001946 | 0.0000222 | 0.0000451 | 0.0000195 | 0.0004494 | 0.0014189 | 0.0001521 | seropositive |
| ABC253 | plate3 | pass | 0.0001676 | 0.0001949 | 0.0000398 | 0.0000560 | 0.0000218 | 0.0002034 | 0.0014829 | 0.0001497 | seropositive |
| ABC094 | plate1 | pass | 0.0002222 | 0.0018643 | 0.0002542 | 0.0000723 | 0.0000201 | 0.0005333 | 0.0002502 | 0.0000980 | seropositive |
| ABC178 | plate2 | pass | 0.0002392 | 0.0018667 | 0.0002508 | 0.0000606 | 0.0000195 | 0.0005059 | 0.0002564 | 0.0000953 | seropositive |
| ABC262 | plate3 | pass | 0.0002389 | 0.0019995 | 0.0002458 | 0.0000700 | 0.0000218 | 0.0002066 | 0.0002516 | 0.0000963 | seropositive |
| ABC095 | plate1 | pass | 0.0001121 | 0.0050100 | 0.0000905 | 0.0000475 | 0.0001849 | 0.0127760 | 0.0104264 | 0.0056060 | seropositive |
| ABC179 | plate2 | pass | 0.0001142 | 0.0048743 | 0.0000772 | 0.0000351 | 0.0001735 | 0.0056178 | 0.0093144 | 0.0054523 | seropositive |
| ABC263 | plate3 | pass | 0.0001175 | 0.0054174 | 0.0000888 | 0.0000470 | 0.0001726 | 0.0017492 | 0.0107118 | 0.0057032 | seropositive |
| ABC096 | plate1 | pass | 0.0001253 | 0.0009660 | 0.0000889 | 0.0000450 | 0.0028297 | 0.0006546 | 0.0015760 | 0.0003976 | seropositive |
| ABC180 | plate2 | pass | 0.0001289 | 0.0009653 | 0.0000755 | 0.0000325 | 0.0026450 | 0.0017733 | 0.0015114 | 0.0003993 | seropositive |
| ABC264 | plate3 | pass | 0.0001318 | 0.0010049 | 0.0000873 | 0.0000447 | 0.0027798 | 0.0009318 | 0.0015840 | 0.0003925 | seropositive |
| ABC086 | plate1 | pass | 0.0000611 | 0.0003321 | 0.0000810 | 0.0000300 | 0.0000230 | 0.0001416 | 0.0002493 | 0.0001302 | seropositive |
| ABC170 | plate2 | pass | 0.0000589 | 0.0003231 | 0.0000671 | 0.0000195 | 0.0000195 | 0.0001310 | 0.0002555 | 0.0001278 | seronegative |
| ABC254 | plate3 | pass | 0.0000636 | 0.0003245 | 0.0000799 | 0.0000309 | 0.0000245 | 0.0001258 | 0.0002507 | 0.0001267 | seropositive |
| ABC087 | plate1 | pass | 0.0000993 | 0.0015979 | 0.0001374 | 0.0000340 | 0.0001356 | 0.0004305 | 0.0003230 | 0.0007757 | seropositive |
| ABC171 | plate2 | pass | 0.0001002 | 0.0016013 | 0.0001273 | 0.0000214 | 0.0001247 | 0.0002020 | 0.0003306 | 0.0007831 | seronegative |
| ABC255 | plate3 | pass | 0.0001038 | 0.0017038 | 0.0001331 | 0.0000345 | 0.0001273 | 0.0002727 | 0.0003263 | 0.0007849 | seropositive |
| ABC088 | plate1 | pass | 0.0000954 | 0.0009660 | 0.0000501 | 0.0031857 | 0.0076618 | 0.0163200 | 0.0007048 | 0.0005235 | seropositive |
| ABC172 | plate2 | pass | 0.0000959 | 0.0009653 | 0.0000347 | 0.0031547 | 0.0070089 | 0.0122238 | 0.0007033 | 0.0005273 | seropositive |
| ABC256 | plate3 | pass | 0.0000997 | 0.0010049 | 0.0000512 | 0.0035665 | 0.0076606 | 0.0130842 | 0.0007129 | 0.0005220 | seropositive |
| ABC089 | plate1 | pass | 0.0010034 | 0.0003814 | 0.0001552 | 0.0000438 | 0.0006923 | 0.0011795 | 0.0013005 | 0.0012309 | seropositive |
| ABC173 | plate2 | pass | 0.0011488 | 0.0003728 | 0.0001462 | 0.0000313 | 0.0006653 | 0.0012405 | 0.0012595 | 0.0012417 | seropositive |
| ABC257 | plate3 | pass | 0.0011392 | 0.0003755 | 0.0001500 | 0.0000435 | 0.0006541 | 0.0016700 | 0.0013091 | 0.0012640 | seropositive |
| ABC090 | plate1 | pass | 0.0200000 | 0.0200000 | 0.0006473 | 0.0048432 | 0.0020459 | 0.0200000 | 0.0039981 | 0.0029554 | seropositive |
| ABC174 | plate2 | pass | 0.0200000 | 0.0200000 | 0.0006544 | 0.0046928 | 0.0019280 | 0.0191706 | 0.0036708 | 0.0029402 | seropositive |
| ABC258 | plate3 | pass | 0.0200000 | 0.0200000 | 0.0006434 | 0.0054411 | 0.0019921 | 0.0200000 | 0.0040151 | 0.0030622 | seropositive |
| ABC091 | plate1 | pass | 0.0001336 | 0.0141035 | 0.0000733 | 0.0000575 | 0.0000950 | 0.0020521 | 0.0043365 | 0.0010674 | seropositive |
| ABC175 | plate2 | pass | 0.0001382 | 0.0125799 | 0.0000590 | 0.0000453 | 0.0000846 | 0.0095263 | 0.0039688 | 0.0010775 | seropositive |
| ABC259 | plate3 | pass | 0.0001409 | 0.0142686 | 0.0000728 | 0.0000562 | 0.0000902 | 0.0022043 | 0.0043583 | 0.0010917 | seropositive |
| ABC092 | plate1 | pass | 0.0002774 | 0.0016671 | 0.0002548 | 0.0000735 | 0.0021836 | 0.0012417 | 0.0011169 | 0.0074239 | seropositive |
| ABC176 | plate2 | pass | 0.0003032 | 0.0016704 | 0.0002515 | 0.0000618 | 0.0020544 | 0.0061144 | 0.0010900 | 0.0071200 | seropositive |
| ABC260 | plate3 | pass | 0.0003011 | 0.0017807 | 0.0002464 | 0.0000711 | 0.0021301 | 0.0003998 | 0.0011257 | 0.0074281 | seropositive |
| ABC093 | plate1 | pass | 0.0001121 | 0.0017912 | 0.0006236 | 0.0000471 | 0.0000570 | 0.0014922 | 0.0080010 | 0.0016269 | seropositive |
| ABC177 | plate2 | pass | 0.0001142 | 0.0017941 | 0.0006305 | 0.0000346 | 0.0000472 | 0.0008549 | 0.0071837 | 0.0016370 | seropositive |
| ABC261 | plate3 | pass | 0.0001175 | 0.0019184 | 0.0006188 | 0.0000466 | 0.0000556 | 0.0017808 | 0.0081404 | 0.0016810 | seropositive |
You can visualise these results using the ggplot2 R package, loaded previously when we ran the library(tidyverse) line.
classifyResults_output %>%
pivot_longer(
-c(SampleID, Plate, QC_total, pred_class_max),
names_to = "Antigen",
values_to = "RAU"
) %>%
ggplot(aes(x = pred_class_max, y = RAU, fill = pred_class_max)) +
geom_boxplot() +
scale_y_log10() +
scale_fill_manual(values = c(seronegative = "#878787", seropositive = "#d6604d")) +
labs(
x = "Classification",
y = "RAU",
fill = "Classification"
) +
facet_grid(~Antigen) +
theme_bw() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))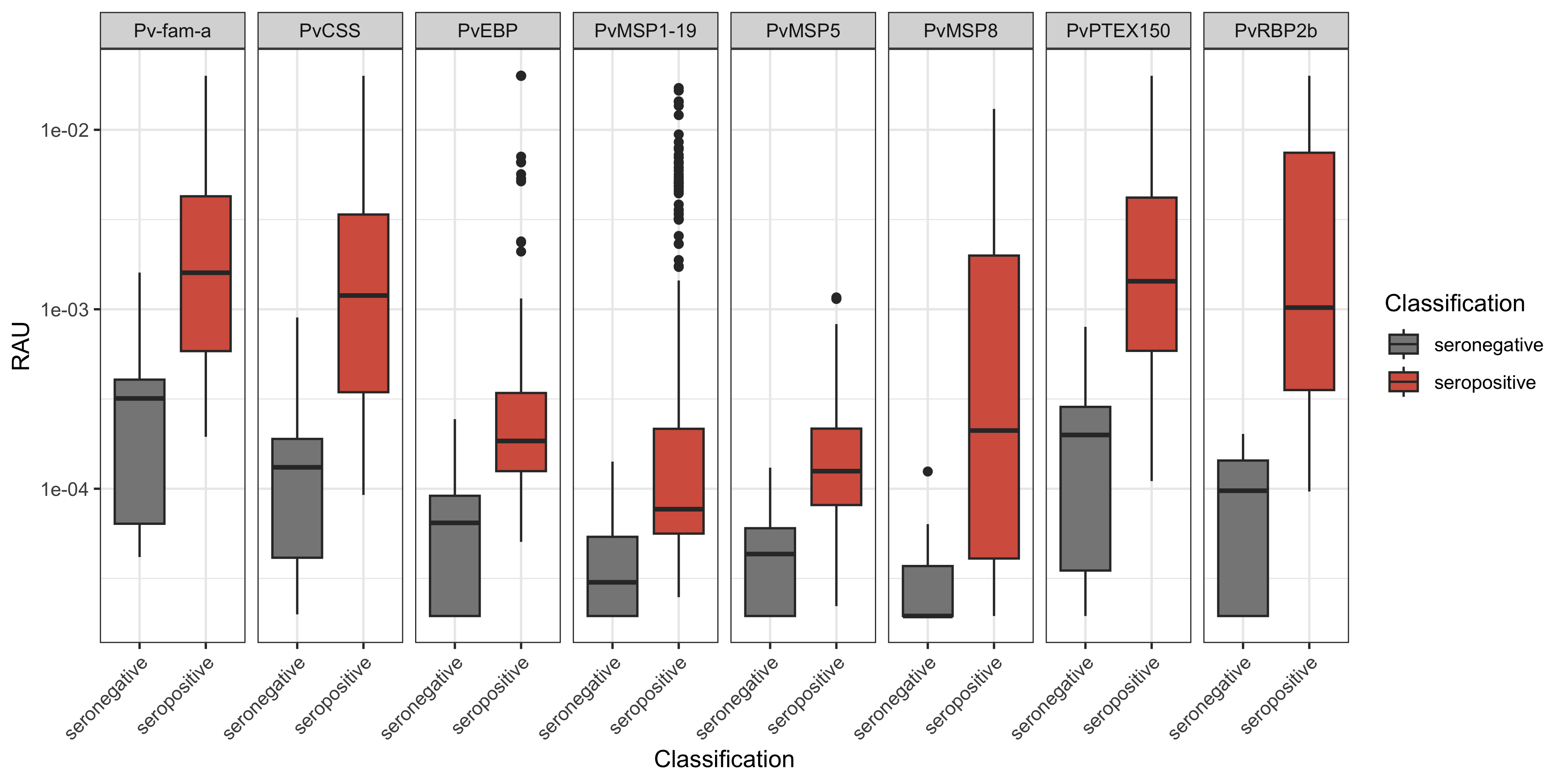
You can also visualise the classification results across multiple sensitivity/specificity thresholds using the plotBoxPlotClassification() function. From here you can choose a specific threshold to visualise.
# Sensitivity/specificity thresholds of interest
sens_spec_all <- c("balanced", "90% sensitivity", "90% specificity")
# Classify results across all thresholds
all_classifications <- purrr::map_dfr(sens_spec_all, ~{
classifyResults(
mfi_to_rau_output = mfi_to_rau_output,
algorithm_type = "antibody_model",
sens_spec = .x,
qc_results = qc_results
) |>
as.data.frame() |>
dplyr::mutate(sens_spec = .x)
})
# Plot classification for a single threshold
plotBoxPlotClassification(all_classifications, "balanced")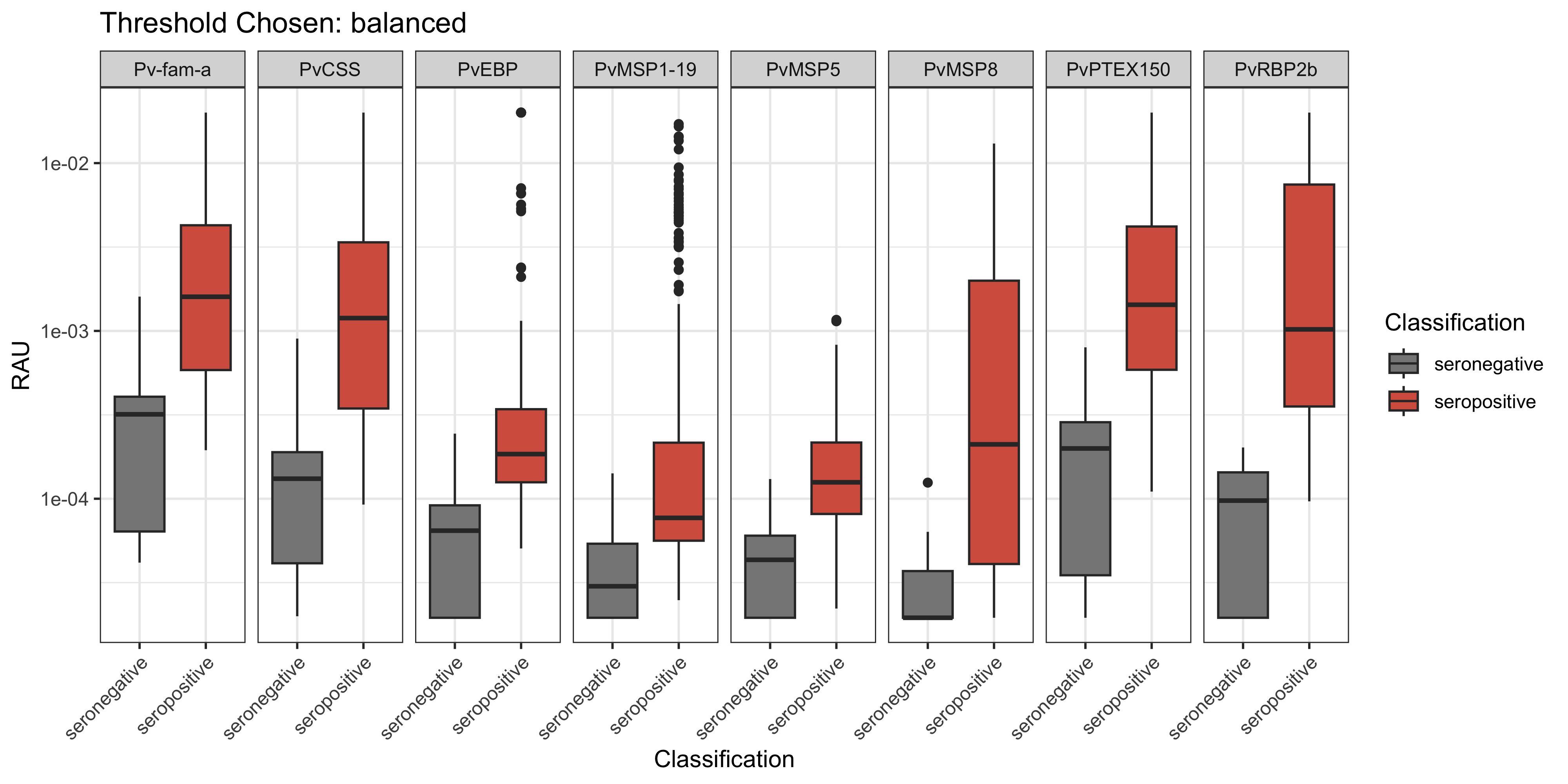
Here are the results of all classification thresholds explored in the PvSeroApp: balanced, 90% sensitivity and 90% specificity. The N’s for the random forest model classifications based on these classifications are displayed.
7.5 Customisations
For more customisations on how to manipulate your ggplot2 see the manual here or the book (Wickham 2016).
